You are here
Morphological data
Work-Package 3 description
Objectives
The objective of this working group is to collect, analyze, and archive morphological descriptions of planktonic communities collected during the Tara Oceans expedition. To do this, various approaches are used:
- Flow- and scan-based technologies for the various size fractions, from viruses to zooplankton
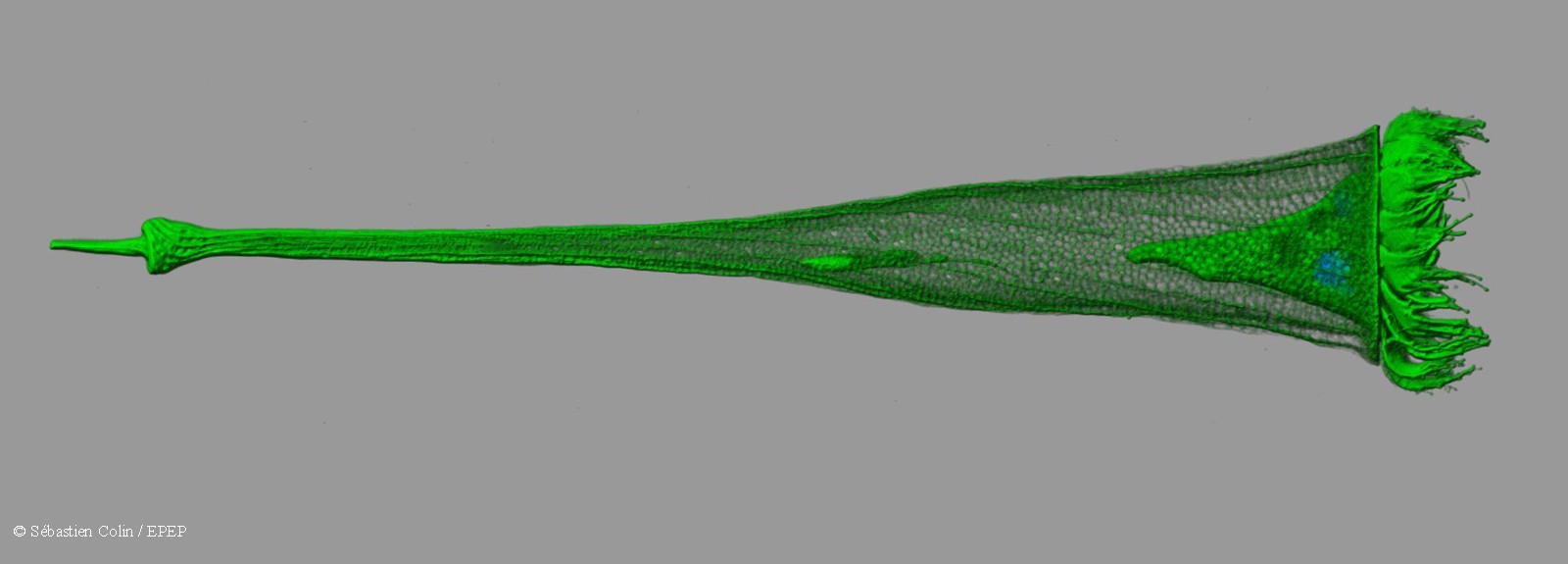
- Development of protocols for high-speed confocal microscopy. 3D reconstructions of organisms (size fraction 5-500μm) are performed using fluorescent markers
- Study of Tara Oceans samples by means of electron microscopy
- Organization of imaging data in various databases in a format compatible with the portal developed by working group no.1 (Archiving samples and data)
This working group is coordinated by Eric Karsenti of EMBL and Colin Sebastian at the Roscoff Biological Station. It involves the UMR 7144 at the Roscoff Biological Station, the Oceanology Laboratory of Villefranche sur Mer, ENS Biology Institute in Paris, and the EMBL in Heidelberg.
Flow- and scan-based technologies - Task 3.1
During each of the stations carried out during the Tara Oceans expedition, the largest plankton and bio-clusters were imaged in situ using a system called UVP (Underwater Vision Profiler) attached to the Niskin-bottle carousel used for sampling in deep waters. Live samples of the size fraction 20-200μm (large protists and small metazoans) were imaged using the onboard FlowCAM system. In addition, fixed or frozen samples were collected to be studied in the lab using the ZooScan system, FlowCAM, or by flow cytometry for the smallest (pico and nano-plankton). This task will provide a collection of related images obtained by all these techniques. The image collection will be used for semi-automatic taxonomic identification of organisms, together with a validation by experts in taxonomy. All data will be integrated into the data warehouse developed in working group no.1 (Archiving samples and data).
High-throughput imaging platform - Task 3.2
In collaboration with Leica Microsystems, EMBL has set up an automated confocal laser scanning system (LCMS) coupled to a computer for recognition of complex phenotypes in cultured cells. This system includes a quick scan of samples at low magnification, an analysis of the samples, followed by a phase of automatic acquisition of images of objects of interest at higher magnification.
OCEANOMICS benefits from this technology for imaging and analysis of protists using fluorescent markers. Fluorescence substantially increases the quality and quantity of biological information in comparison with automated optical microscopy techniques. Several fluorescent markers were used successfully in the Tara Oceans expedition to mark the membrane, microtubules, nuclei, cell walls, lipids, etc.
Based on this system developed at EMBL, an imaging procedure in 2 stages has been developed by the OCEANOMICS team. The first step is to identify objects of interest during a quick scan at low magnification, coupled with automated analysis. The objects are then imaged in 3 dimensions at higher magnification, better resolution, and using more markers.
Following promising tests, this methodology has been improved to increase the speed of acquisition in relation to the number of samples to be analyzed, to enable broad coverage of size fractions to be studied (1-500μm) and to improve the classification procedure of objects characterized by complex diversity.
The vast collection of generated images will be stored in the data warehouse developed by working group no.1 and be made accessible to the greatest possible number of researchers.
Electron microscopy - Task 3.3
Electron microscopy provides important information, relatively unknown until now, about the cellular organization of planktonic organisms.
Scanning electron microscopy was used to reveal the wide variety of planktonic exoskeletons made of silica, limestone, or chitin. Transmission electron microscopy, in turn, allows the study of subcellular structures such as organelles, vesicles and cell walls, but also the associations between organisms (host-pathogen, or host-symbiont interactions).
Obtaining such information, both qualitative and quantitative, is critical for understanding the functioning of planktonic ecosystems and their biogeochemical status, particularly regarding eukaryotes.
During the Tara Oceans expedition, organisms from size fractions 5-20μm and 20-200μm were fixed in order to apply these 2 approaches. In the context of OCEANOMICS, these samples are analyzed using routine protocols. Ultimately, electron microscopy data will be useful to interpret and validate the phenotypic hypotheses developed on genetic data generated by working group no.4 (Production and primary analysis of genetic data).