You are here
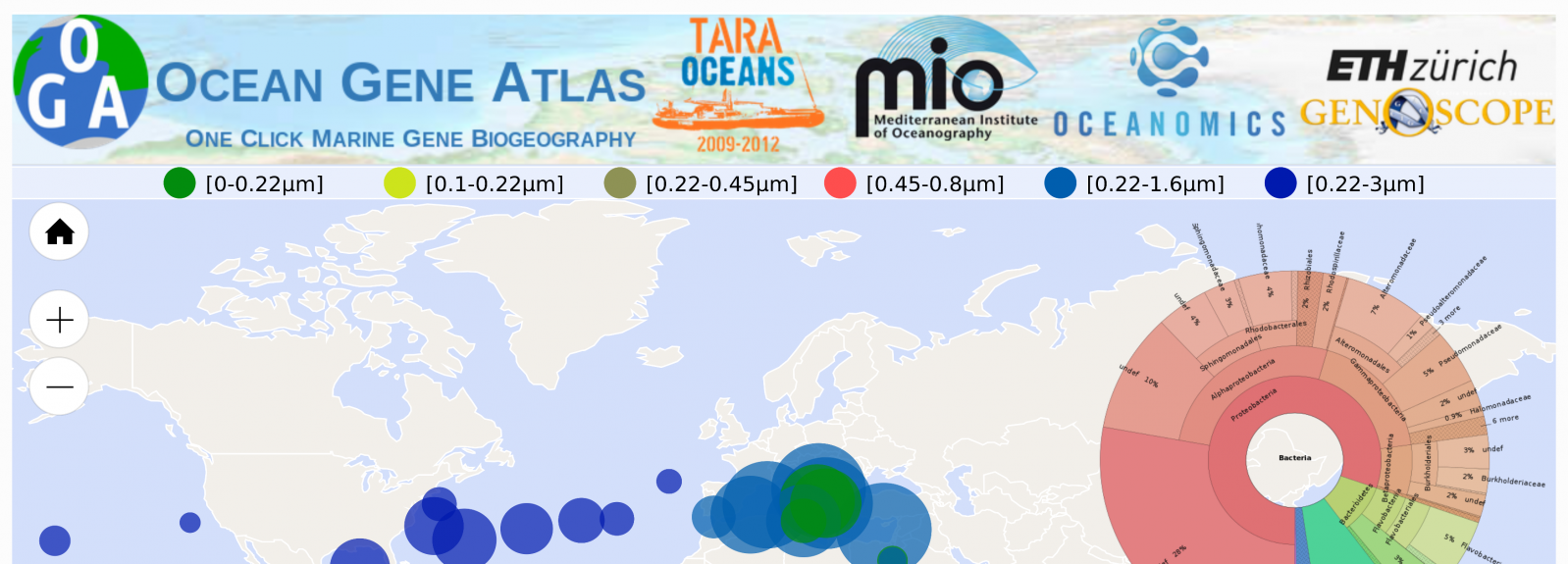
Ocean Gene Atlas, explore marine Big Data with nothing but a web browser
Researchers from the Mediterranean Institute of Oceanology (Marseille) and the Roscoff Biological Station developed a website to easily query the recently published Tara-Oceans genomic resources.
Ocean Gene Atlas is a freely-available web service at:
http://tara-oceans.mio.osupytheas.fr/ocean-gene-atlas/
Gigantic datasets, treasure troves for functional ecogenomics
To understand the mechanisms by which genes influence phenotype in the wild, biologists frequently apply reductionist approaches that target specific marker genes known or hypothesized to play a role in processes such as metabolic pathways, biotic and abiotic interactions. Testing such candidate marker gene hypotheses requires detailed analyses of voluminous genomic data in their precise environmental context, a task which requires extensive expertise in interrogating heterogeneous data types (i.e. sequencing reads, assemblies, genes together with their taxonomic and functional annotations, environmental variables) to provide integrated interpretations.
Recently, the Tara Oceans pan-oceanic expedition deployed a holistic sampling followed by deep omics sequencing of plankton ranging in size from viruses to fish larvae, coupled with comprehensive in situ biogeochemical measurements which provide the detailed environmental contexts necessary for ecological interpretation of marine microbiomes.
The first release of Ocean Gene Atlas (OGA) empowers users to search by means of gene similarity the two largest marine gene catalogs ever released to date (both produced by the Tara Oceans consortium, which combined represent all three eukaryotic, prokaryotic and viral lineages). With just one click, the abundance and location of target sequences are visualized on world maps as well as their taxonomic distribution, all in the context of detailed environmental features (temperature, nutrients, etc.) measured at the time of sampling.
No informatic skills needed
By hiding the complex and time consuming integration of heterogeneous data sources behind a user-friendly minimalist web form, the OGA web server aims to broaden the access to the rapidly accumulating environmental marine genomics datasets. Enabling marine biologists to mine such data - without specific high performance hardware or programming skills - is key to extract knowledge from these valuable but still underexploited resources.
To be continued!
The prerequisites for inclusion in OGA are the availability of the three core resources: gene sequence catalogs, gene abundance estimates in biosamples, and geolocalized environmental context of biosamples. The OGA catalog collection will be periodically updated as further marine environmental genomics datasets are publically released (from Tara Oceans but also other marine biome projects). Future plans also include addition of modules to improve results interpretation, most notably to refine taxonomical analysis by on the fly building of phylogenetic trees as well as additional graphical representations of environmental data.
Reference in Nucleic Acids Research:
Emilie Villar, Thomas Vannier, Caroline Vernette, Magali Lescot, Miguelangel Cuenca, Aurélien Alexandre, Paul Bachelerie, Thomas Rosnet, Eric Pelletier, Shinichi Sunagawa, Pascal Hingamp. (2018) The Ocean Gene Atlas: exploring the biogeography of plankton genes online, Nucleic Acids Research, gky376, http://doi.org/10.1093/nar/gky376
Contacts:
- Emilie Villar, Sorbonne Universités, UPMC Université Paris 06, CNRS, Laboratoire Adaptation et Diversité en Milieu Marin UMR7144, Station Biologique de Roscoff, Roscoff, France,
- Pascal Hingamp, Aix Marseille Univ, Université de Toulon, CNRS, IRD, MIO UM 110, 13288, Marseille, France, , Tel: +33 4 86 09 06 66; Fax: +33 4 86 09 06 41